v.1.0.7
Launch
Optimize bacterial or archaeal genome bins using a dereplication, aggregation and scoring strategy
DAS Tool is an automated method that integrates the results of a flexible number of binning algorithms to calculate an optimized, non-redundant set of bins from a single assembly.
Implemented for KBase by Sean Jungbluth([email protected])
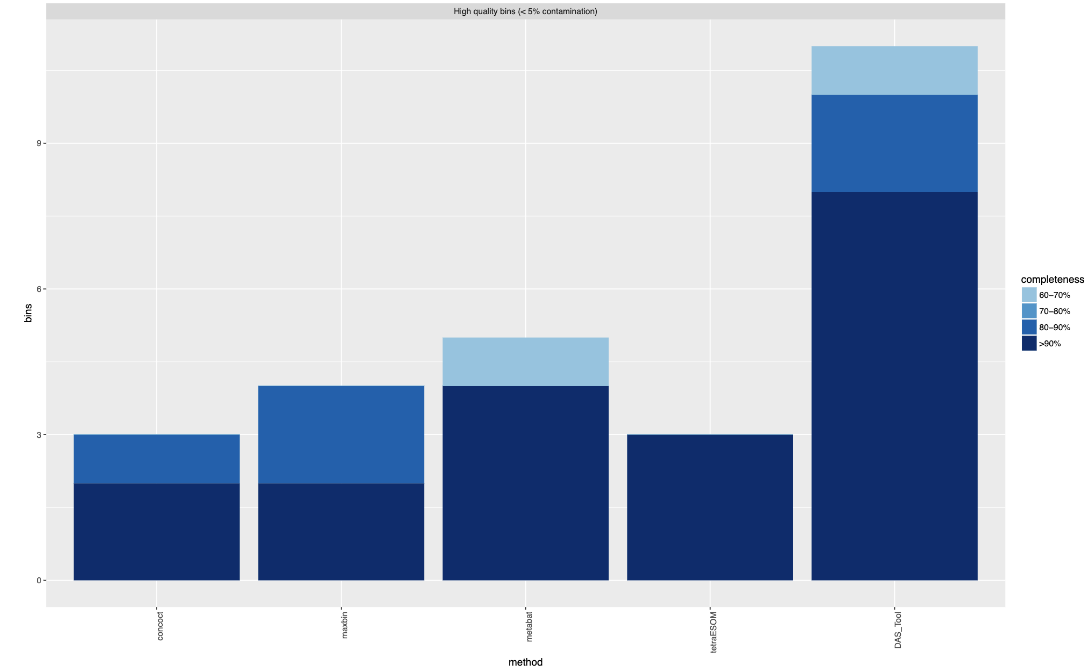
DAS Tool takes metagenome "bins" generated from the same assembly, but using different genome binning tools, and optimizes their content using a dereplication, aggregation and scoring strategy.
Configuration:
Assembly Object: The Assembly object is a collection of assembled genome fragments, called "contigs". These are part of the input file required for DAS Tool to run. Currently only a single Assembly object is accepted by the DAS Tool App.
Input BinnedContig Object Names: The BinnedContig Objects represents the different genome binning tool output binned contigs to be used for input to DAS Tool.
Output BinnedContig Object Name: The output BinnedContig Object is produced by DAS Tool.
Output:
Output Object:The BinnedContig Object represents the directory of binned contigs created by DAS Tool. This object can be used for downstream analysis.
Output Bin Summary Report:The number of bins produced, the number of contigs that were binned and the total number of contigs in the assembly.
Downloadable files: The enitre output of the DAS Tool run may be downloaded as a zip file.
Related Publications
- Sieber CMK, Probst AJ, Sharrar A, Thomas BC, Hess M, Tringe SG, Banfield JF. Recovery of genomes from metagenomes via a dereplication, aggregation and scoring strategy. 2018; 3(7): 836-843. doi:10.1038/s41564-018-0171-1 , https://doi.org/10.1038/s41564-018-0171-1
- DAS_Tool source: , https://github.com/cmks/DAS_Tool
- Hyatt D, Chen G-L, LoCascio PF, Land ML, Larimer FW, Hauser LJ. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics. 2010;11: 119. doi:10.1186/1471-2105-11-119 , https://www.ncbi.nlm.nih.gov/pubmed/20211023
- Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL. BLAST+: architecture and applications. BMC Bioinformatics. 2009;10: 421. doi:10.1186/1471-2105-10-421 , https://www.ncbi.nlm.nih.gov/pubmed/20211023
- Buchfink B, Xie C, Huson DH. Fast and sensitive protein alignment using DIAMOND. Nature Methods. 2015;12: 59-60. doi:10.1038/nmeth.3176 , https://www.ncbi.nlm.nih.gov/pubmed/25402007
- Pullseq: , https://github.com/bcthomas/pullseq
- R: A Language and Environment for Statistical Computing: , http://www.R-project.org/
- Ruby: A Programmers Best Friend: , https://www.ruby-lang.org
App Specification:
https://github.com/kbaseapps/kb_das_tool/tree/c70fdbdef91d5b6614dfd387fa127c1492769a8f/ui/narrative/methods/run_kb_das_toolModule Commit: c70fdbdef91d5b6614dfd387fa127c1492769a8f